Wisconsin Epidemiological Study of Diabetic Retinopathy (WESDR)
is an epidemiological study of a cohort of diabetic
patients receiving their medical care in an 11-county area in Southern
Wisconsin. Detailed descriptions of the data can be found in
Klein et al. (1988). A number of medical, demographic, ocular and
other covariates were recorded at the baseline and later examinations
along with a retinopathy score for each eye. As in Wahba et al. (1995),
we investigate how progression of diabetic retinopathy at the first
follow-up depends on the following covariates:
dur (duration of diabetes at baseline),
gly (glycosylated hemoglobin, a measure of hyperglycemia), and
bmi (body mass index = weight in kg/(height in m)![]() ).
As in Wahba et al
).
As in Wahba et al ![]() , we chose a subgroup of the younger onset
consisting of 669 subjects with no or non-proliferative
retinopathy at the baseline. See Wahba et al. (1995) for details of
this data set.
, we chose a subgroup of the younger onset
consisting of 669 subjects with no or non-proliferative
retinopathy at the baseline. See Wahba et al. (1995) for details of
this data set.
We use this data set to illustrate how to fit smoothing
spline models for non-Gaussian data. Firstly, we fit a simple cubic spline
> wesdr.fit1<- ssr(prg~bmi, rk=cubic(bmi), data=wesdr, family="binary",
scale=T, spar="u", varht=1)
> summary(wesdr.fit1)
...
Coefficients (d):
(Intercept) bmi
-1.286666 1.793480
UBR estimate(s) of smoothing parameter(s) : 8.793603e-06
Equivalent Degrees of Freedom (DF): 6.036309
Estimate of sigma: 1
> grid <- data.frame(bmi=seq(min(wesdr$bmi),max(wesdr$bmi),len=100))
> p.wesdr.fit1 <- predict(wesdr.fit1, grid)
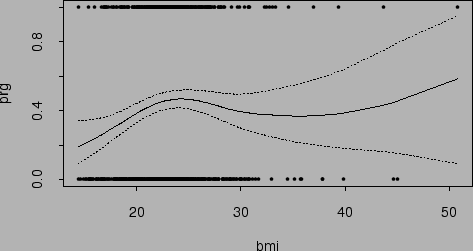
Figure ![]() shows the fitted probability function and
its 95% Bayesian confidence intervals based on wesdr.fit1.
shows the fitted probability function and
its 95% Bayesian confidence intervals based on wesdr.fit1.
 |
Wahba et al ![]() reached the following model
reached the following model
wesdr.fit2<- update(wesdr.fit1, prg~dur+gly+bmi+I(dur*bmi),
rk=list(cubic(dur), cubic(bmi), rk.prod(kron(dur), cubic(bmi)),
rk.prod(kron(bmi), cubic(dur)), rk.prod(cubic(dur), cubic(bmi))))
> summary(wesdr.fit2)
...
Coefficients (d):
(Intercept) dur gly bmi I(dur * bmi)
-6.1810142 -2.5728071 0.3864040 0.1401764 11.2939414
UBR estimate(s) of smoothing parameter(s) : 2.653578e+00 3.982479e+00 2.941409e+05 3.086313e+05 8.194281e-02
Equivalent Degrees of Freedom (DF): 11.19937
Estimate of sigma: 1
> grid <- expand.grid(dur=seq(min(wesdr$dur),max(wesdr$dur),len=40),
bmi=seq(min(wesdr$bmi),max(wesdr$bmi),len=40),
gly=median(wesdr$gly))
> p.wesdr.fit2 <- predict(wesdr.fit2, grid)
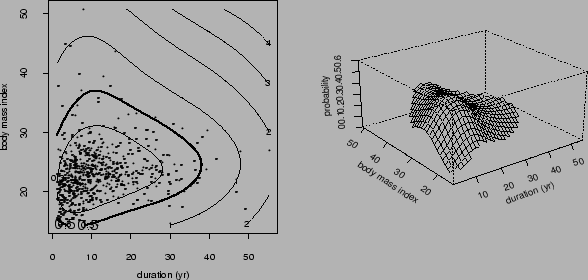
Figure ![]() reproduce Figure 6.1 in Wahba et al (1994).
reproduce Figure 6.1 in Wahba et al (1994).
 |